Last modified 27th April '95 © Birkbeck College 1995
Back to main PPS Index
Back to Quaternary Structure Index
Multienzyme Complexes
In a number of metabolic pathways, several enzymes which catalyze different
stages of the process have been found to be associated noncovalently, giving
a multienzyme complex. The proximity of the different types of
enzymes increases the efficiency of the pathway: the overall reaction rate is
increased with respect to catalysis by unassociated units, and side reactions
are minimized. In some cases molecular mechanisms have been identified
for the transfer of metabolites from one enzyme to the next within the complex.
Refer to
Simon Brocklehurst's Multienzyme Complexes page, and to the
enzymes section in a previous chapter.
Pyruvate Dehydrogenase Complex
This multienzyme complex catalyses the conversion of pyruvate and
coenzyme A (CoA) to acetyl CoA.
The reaction
There are four stages in this
pathway, which are catalyzed by three enzymes:
- "E1" - pyruvate dehydrogenase
- This enzyme catalyzes the decarboxylation of pyruvate. This involves the
prosthetic group thiamine pyrophosphate, or TPP.
- "E2" - dihydrolipoyl transacetylase
- Two steps of the pathway are catalyzed by this enzyme:
- oxidation of the 2-carbon (acetyl) unit, which is transferred to the
lipoamide prosthetic group of the enzyme, giving an acetyllipoamide
group
- transfer of the acetyl group from the lipoamide to CoA, giving acetyl CoA
- "E3" - dihydrolipoyl dehydrogenase
- Finally, this enzyme regenerates the oxidized form of lipoamide. This involves
the FAD prosthetic group.
Note that TPP, lipoamide and FAD are catalytic cofactors which remain
unaltered by the net reaction, whereas CoA and NAD+ are stoichiometric
cofactors; the overall reaction is:
pyruvate + CoA + NAD+ ----> acetyl CoA + carbon dioxide + NADH
The four stages are summarized in this diagram .
Note that the lipoamide cofactor of E2 reacts with the product
(hydroxyethyl-TPP) from E1, and (in its modified form,
dihydrolipoamide, formed by the third step) interacts with E3.
It is for this reason that the interaction rate is increased by proximity of
all three enzymes, and in fact the lipoamide group is a long flexible arm about
14Å long. It is
covalently bonded to a specific lysine residue
of the enzyme. The movement of the arm may be driven by a change in the net charge
of the lipoyl group, depending on the ionization of the sulphydryl groups.
The structure of the complex
In isolation, E2 forms a homo-24-mer, believed to have cubic symmetry.
The E1 and E3 subunits each form homodimers. Electron micrographs
indicate that the whole complex has a cubic arrangment, in which an E3
dimer associates with each face of the E2 24-mer cube, while an E1
dimer is positioned on each edge of the cube.
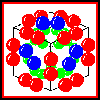 Click here for a diagram
of this model.
Click here for a diagram
of this model.
Refer to Reed(1974).
The Electron Transport Chain
Oxidative phosphorylation occurs in the mitochondria. The various components
of the electron transport chain are situated in the inner membrane of this organelle.
Oxidation of NADH and FADH2 (produced by the citric acid cycle) results in free energy which is coupled to the
phosphorylation of ADP to form ATP (by ATP synthase, another complex in the
inner mitochondrial membrane). Electrons pass from lower to higher standard
reduction potentials in the chain in a series of redox reactions;
the ultimate electron acceptor is oxygen (O2).
The electron transport chain in fact consists of four multienzyme complexes. These complexes
are free to diffuse laterally through the membrane, and are not present in the
same numbers.
- Complex I: NADH - Coenzyme Q reductase, 26 subunits, 850 kD.
This large protein includes one molecule of the prosthetic group flavin
mononucleotide , or FMN, and six to seven iron-sulphur clusters
- Complex II: succinate - Coenzyme Q reductase, 5 subunits, 127 kD. Includes:
- FAD, covalently bound
- three iron-sulphur clusters
- cytochrome b560
- Complex III: Coenzyme Q - cytochrome c reductase, 10 subunits, 280kD
- Complex IV: cytochrome c oxidase 7-8 subunits, 160-170kD
- cytochrome a
- cytochrome a3
Other components of the chain are succinate (between complexes I and II in the
sequence), Coenzyme Q (between II and III) and
cytochrome c
(between III and IV).
Electron micrographs have indicated the overall shape of Complex IV and its
orientation relative to the inner mitochondrial membrane. Techniques involving
chemical cross-linking and antibody-binding to different subunits have led to
a model of the whole complex, shown in this diagram
(after Brunori and Wilson, 1982).The cytochrome c oxidase complex
catalyzes the oxidation of four reduced cytochrome c molecules; the
binding of cytochrome c to the complex mainly involves subunit II as
indicated.
References
Pyruvate Dehydrogenase Complex
- Reed, L.J., (1974) Multienzyme complexes, Acc. Chem. Res. 7,
40-46
- Stryer, L., (1981) Biochemistry, W.H. Freeman & Co., New York pp. 290-294
- Voet, D. and Voet, J.G. (1990) Biochemistry, John Wiley & Sons, New York, pp. 186-188
Electron Transport Chain
- Brunori, M. and Wilson, M.T. (1982) Cytochrome oxidase, Trends Biochem. Sci.
7, 295-299
- Capaldi, R.A. (1982) Arrangement of proteins in the mitochondrial inner
membrane, Biochim. Biophys. Acta 695, 291-306
- Capaldi, R.A., Malatesta, F. and Darley-Usmar, V.M. (1983) Structure of
cytochrome c oxidase, Biochim. Biophys. Acta 726, 135-148
- Darnell, J., Lodish, H. and Baltimore, D. (1986) Molecular Cell Biology,
W.H. Freeman & Co., New York pp. 873-889
- Poulos, T.L. and Kraut, J. (1980) A hypothetical model of the cytochrome
c peroxidase . cytochrome c electron transfer complex,
J. Biol. Chem. 255 10322-10330
- Scott, R.A. (1989) X-ray absorption spectroscopic investigations of
cytochrome c oxidase structure and function, Annu. Rev. Biophys.
Biophys. Chem. 18 137-158
- Stryer, L., (1981) Biochemistry, W.H. Freeman & Co., New York pp. 319-320
- Sweeney, W.V. and Rabinowitz, J.C. (1980) Proteins containing 4Fe-4S clusters:
an overview, Annu. Rev. Biochem. 49, 139-161
- Voet, D. and Voet, J.G. (1990) Biochemistry,
John Wiley & Sons, New York, pp. 532-544
Back to Main PPS Index
J. Walshaw
 Click here for a diagram
of this model.
Click here for a diagram
of this model.
 Click here for a diagram
of this model.
Click here for a diagram
of this model.