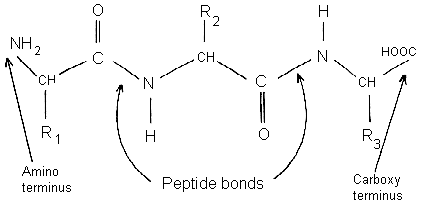

Amino acids in proteins (or polypeptides) are joined together by peptide bonds.
The sequence of R-groups along the chain is called the primary structure. Secondary structure refers to the local folding of the polypeptide chain. Tertiary structure is the arrangement of secondary structure elements in 3 dimensions and quaternary structure describes the arrangement of a protein's subunits.
Linus Pauling and Robert Corey analysed the geometry and dimensions of the peptide bonds in the crystal structures of molecules containing one or a few peptide bonds. Their results are summarised in this diagram where the consensus bond lengths are shown in Angstrom units. Bond angles in degrees are also shown for the peptide N and C atoms.
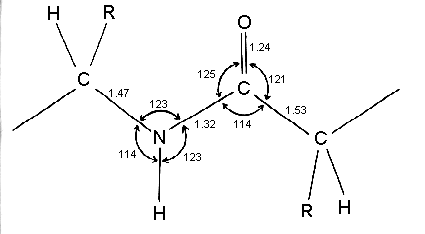
Note that the C-N bond length of the peptide is 10% shorter than that found in usual C-N amine bonds. This is because the peptide bond has some double bond character (40%) due to resonance which occurs with amides. The two canonical structures are:
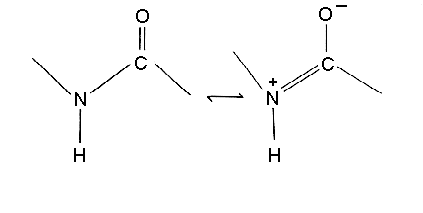
As a consequence of this resonance all peptide bonds in protein structures are found to be almost planar, ie atoms Calpha(i), C(i), O(i), N(i+1) H(i+1) and Calpha(i+1) are approximately co-planar. This rigidity of the peptide bond reduces the degrees of freedom of the polypeptide during folding.
The peptide bond nearly always has the trans configuration since it is more favourable than cis, which is sometimes found to occur with proline residues.
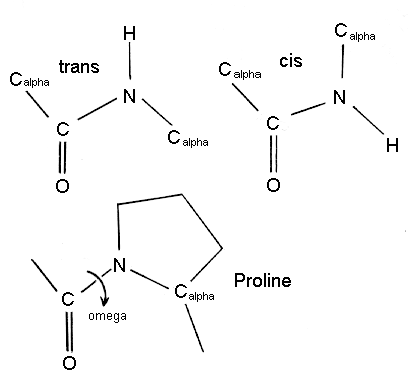
As can be seen above, steric hindrance between the functional groups attached to the Calpha atoms will be greater in the cis configuration. However for proline residues, the cyclic nature of the side chain means that both cis and trans configurations have more equivalent energies. Thus proline is found in the cis configuration more frequently than other amino acids. The omega torsion angle of proline will be close to zero degrees for the cis configuration, or most often, 180 degrees for the trans configuration.