 Three Dimensional NMR Spectroscopy
Three Dimensional NMR Spectroscopy Three Dimensional NMR Spectroscopy
Three Dimensional NMR Spectroscopy
A three dimensional NMR experiment (see picture above) can easily be constructed from a two dimensional one by inserting an additional indirect evolution time and a second mixing period between the first mixing period and the direct data acqusition. Each of the different indirect time periods (t1, t2) is incremented separately. There are two principal classes of 3D experiments: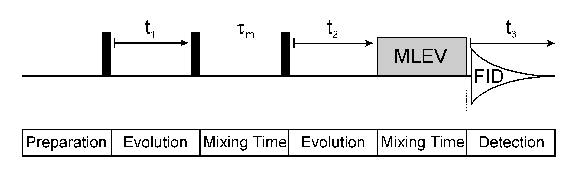
NOESY-HSQC, TOCSY-HSQC:
2D spectra (like
NOESY or
TOCSY) of larger proteins are often
crowded with signals. Therefore, these spectra are spreading out
in a third dimension (usually 15N or 13C),
so that the signals are distributed in a
cube instead of a plane. This spreadout is achieved by combining
HSQC
and NOESY in a single 3D experiment: The NOESY experiment
is extended by an HSQC step. Acquisition starts after this HSQC
step instead of at the end of the NOESY mixing time. The
resulting experiment is called 3D NOESY-HSQC.
In a similar way, a TOCSY-HSQC can
be constructed by combining the TOCSY and the HSQC experiment.
The experiments HCCH-TOCSY and HCCH-COSY are alternatives to the 13C-TOCSY-HSQC experiment which shows a markedly reduced sensitivity for larger proteins. In both experiments the magnetization is transferred via direct 1J couplings between the atoms, allowing a much faster magnetization transfer as in the TOCSY-HSQC. Magnetization is transferred (blue arrows) from a sidechain (or backbone) proton (red) to the direct attached carbon atom (yellow), by 1J coupling to the neighboring carbon atoms (yellow) and finally to their attached protons (red). Therefore, a HCCH-TOCSY experiment looks in principle the same like a 13C-TOCSY-HSQC. The type of an amino acid can be recognized from the peak pattern in the usual way, as can be seen from the picture below. It shows some regions from different planes of a HCCH-TOCSY which together contain the complete spin system of a leucine residue: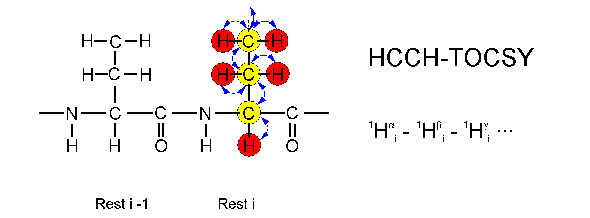
The most important advantage of the triple resonance spectra is their simplicity: They contain only a few signals on each frequency - often only one. The problem of spectral overlap is therefore markedly reduced (this is the main reason, why proteins of more than 20 kDa can be assigned with triple resonance experiments). However, the coordinates of clearly separated signals from different amino acids can accidentally be identical ('degeneration' of signals). The correct choice of connectivities between amino acids is the main problem in the assignment of triple resonance spectra.
Another advantage of triple resonance spectra is their high sensitivity which is caused by an efficient transfer of magnetization. The magnetization is tranferred via 1J or 2J couplings (i.e. directly via the covalent chemical bonds). Therefore, the transfer times are shorter and the losses due to relaxation are smaller than in homonuclear experiments.
The disadvantage of all triple resonance experiments is the necessity of doubly labelled proteins, the preparation of which is often expensive.
There is a whole bunch of triple resonance experiments which can not be covered in this short introduction. Therefore, I will explain only the general nomenclature of triple resonance experiments and I will deal with the HNCA which is the prototype for all these experiments.
If you like to learn more about the various triple resonance experiments (and of all the other nice NMR methods), just have a look at the database of NMR experiments at the Universidad Autonoma de Barcelona.
In the HNCO experiment the magnetization is
transferred (blue arrows) from the
HN(i) proton via the N(i) atom to the directly attached CO(i-1)
carbon atom and return the same way to the HN(i) nucleus which
is directly detected. The frequencies of all three nuclei (red) are
detected.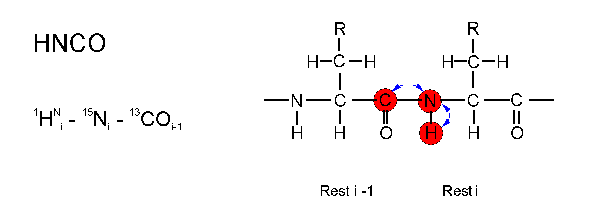
In the HN(CA)CO experiment
the magnetization is transferred from
the HN(i) proton via the N(i) atom and the CA nucleus
(Calpha(i)) to the CO(i) carbon atom and back the same way. The
Calpha atom (yellow) acts only as relay nucleus, its frequency is not
detected. It is only the frequencies of HN, N and CO (red) which are
detected.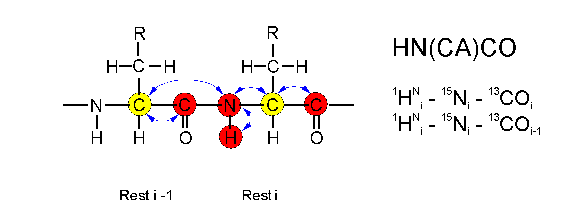
This nomenclature has the advantage, that the spectra can be
easily imagined by their names:
In HNCO an amide proton is correlated with the CO atom of the
preceeding amino acid, whereas in HN(CA)CO the correlation to
the intraresidual CO atom is also visible.
The HNCA experiment is the prototype of all triple resonance experiments. Starting at an amide proton (H) the magnetization is transferred to the directly attached nitrogen atom (N) which is measured as the first spectral dimension. Then the magnetization is transferred to the Calpha nucleus (CA) which is measured as second dimension. Afterwards, the magnetization is transferred back the same way to the amide proton which is measured as the third (direct) dimension.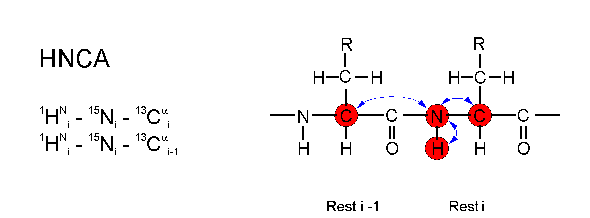
In each step magnetization is transferred via strong 1J couplings between the nuclei. The coupling which connects the nitrogen atom with the Calpha carbon of the preceeding amino acid (2J= 7 Hz) is only marginally smaller than the coupling to the directly attached Calpha atom (1J = 11 Hz). Thus, the nitrogen atom of a given amino acid is correlated with both Calpha - its own and the one of the preceeding amino acid.
Therefore, it is possible to assign the protein backbone exclusively with an HNCA spectrum. But usually more triple resonance experiments are needed because the cross signal of the preceeding amino acid has to be identified and degenerate resonance frequencies have to be resolved (see assignment chapter).