 Transport of Proteins Across Membranes
Transport of Proteins Across Membranes
 Index to Course Material
Index to Course Material
 Index to Section 4
Index to Section 4
(This page has links to sections of Mark Dalton's
Introduction
to Cell Biology; the
Cell Biology Notes at MIT will also be of use.)
Some proteins function in the cytoplasm of cells, while others are an integral
component of membranes. Others are
directed towards particular organelles, while a number of types are secreted
(exported from the cell). Most proteins which are not cytoplasmic enter the
lumen of the Endoplasmic Reticulum (E.R.) as they are synthesized.
Endoplasmic Reticulum
The Endoplasmic Reticulum (E.R.) is an extensive network of flattened,
membrane-bound sacs generally arranged in layers in the cell. Some E.R. has
ribosomes associated with the outer (cytosolic) side of its membrane, and is
called rough endoplasmic reticulum due to its studded appearance.
Refer to the Mark Dalton's section on
the
endoplasmic reticulum, and the section on
Structure and Function of Organelles (including a diagram of
the endoplasmic
reticulum) in the
Cell Biology Notes
at MIT.
The polypeptide chains synthesized by the ribosomes bound to the rough E.R.
enter the endoplasmic reticulum lumen. There is no difference between the
ribosomes associated with E.R. and those which are free in the cytosol:
- ribosomes isolated from the rough E.R. are found to be able to synthesize
cytosolic proteins from their mRNA sequences
- ribosomes obtained from the cytosol, when supplied with the appropriate
mRNA and rough E.R. membranes which have been stripped of their ribosomes, are able
to synthesize secretory and other proteins which normally enter the E.R.
Therefore whether or not a ribosome binds to the E.R. is dictated by the type
of protein which it is synthesizing.
The ribosome associates with the E.R. membrane by means of a protrusion of the
large (60S) subunit into the lipid bilayer.
Signal peptides
Proteins which enter the E.R. during synthesis, and which are therefore
synthesized by ribosomes on the rough E.R., have a signal peptide of
16 to 30 residues at their N-terminus. This sequence is subsequently cleaved
off (by signal peptidase).
The signal peptides of different proteins are not homologous, but they
generally include several (4 to 12) hydrophobic residues, which are found to
be essential for the polypeptide to cross the E.R. membrane. There is also a
basic residue a few residues before the hydrophobic sequence.
Some examples of signal peptides (continuous hydrophobic sequences are in bold):
Preproalbumin M K W V T F L L L L F I S G S A F S
Pre-IgG light chain M D M R A P A Q I F G F L L L L F P G T R C
Prelysozyme M R S L L I L V L C F L P L A A L G
Preprolactin M N S Q V S A R K A G T L L L L M M S N L
Prepenicillinase M S I Q H F R V A L I P F F A F C L P V F A
Prevesicular stomatitis M K C L L Y L A F L F I H V N C
virus glycoprotein
Prelipoprotein M K A T K L V L G A V I L G S T L L A G
Genetic engineering techniques can add a signal peptide to the N-terminus of
cytoplasmic proteins, such as globin,which naturally have no such sequences;
this results in the engineered protein entering the E.R. and having the
signal sequence cleaved off.
Cotranslation transport
In cell-free systems containing ribosomes but no E.R. membrane component, the
mRNA of secretory protein sequences is translated to give the polypeptide
complete with signal sequence, which is not cleaved. If membrane vesicles are
then added to the system, the protein still does not enter them, and the
signal peptide remains attached (in the majority of proteins; see
below).
The transport of most proteins through the membrane is therefore
cotranslational. However, membrane vesicles need not be present at the
beginning of translation. In the system described above, addition of membrane
before the approximately the first 70 residues have been polymerized is
sufficient for normal transport and signal cleavage to occur.
Posttranslational transport
There are exceptions to the cotranslational behaviour described above.
Mitochondria and chloroplasts are organelles which contain very small amounts
of DNA. Most of their proteins are however encoded by the nuclear genome, and
are synthesized by free cytoplasmic ribsomes. The resulting polypeptides have
N-terminal sequences which are removed after the protein has crossed the
organelle membrane. Such sequences are clearly of a different nature to those
which associate their translating ribosomes with the E.R. However, some yeast
secretory proteins have been found to be capable of passing through the E.R.
after translation is complete, but the process requires ATP.
Uncleaved signal sequences
 Ovalbumin 1ova (1.0Mb) [Bbk|BNL|ExP|Waw|Hal]
is an example of a secretory protein which does not naturally have its
signal sequence cleaved. The 100 N-terminal residues are found to be necessary
for transport through the membrane to be effected.
Ovalbumin 1ova (1.0Mb) [Bbk|BNL|ExP|Waw|Hal]
is an example of a secretory protein which does not naturally have its
signal sequence cleaved. The 100 N-terminal residues are found to be necessary
for transport through the membrane to be effected.
Signal Recognition Particles
Secretory and other non-cytoplasmic proteins are synthesized only in association
with the E.R., and not with any other kind of membrane. Therefore the E.R. membrane
must have some kind of identifying feature. The signal recognition particle (SRP)
consists of six polypeptide chains and a 300-nucleotide RNA. This was identified
by stripping E.R. membrane of certain membrane proteins, without which the
membrane cannot accommodate the secretory polypeptide chains.
In the absence of E.R. membrane vesicles, but with SRP present, the synthesis
of secretory proteins is halted when the polypeptide is approximately 70 amino
acids long. Addition of membrane vesicles allows translation to resume. The
interaction of the ribosome/polypeptide/SRP complex with the membrane is
mediated by SRP receptor, a 650-residue integral membrane protein which
may bind to the ribosome as well as to SRP.
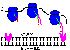 Here is a diagram.
Here is a diagram.
Prokaryotic Signal Sequences
Prokaryotic cells have no organelles such as E.R., but they do have ribosomes
bound to the plasma membrane which synthesize secreted proteins, such as
maltodextrin-binding
protein which is secreted into the space between the
plasma membrane and the cell wall (the periplasmic space) in gram
negative bacteria. Such secreted proteins have similar N-terminal peptide
sequences to eukaryotic secreted proteins, which are cleaved following
secretion. Genetically engineered varieties of maltose-binding protein, in which
only a single hydrophobic residue of the signal peptide has been replaced by a
charged one, are not secreted, but remain in the cytoplasm complete with
attached signal sequence.
Other examples, all with a similar type of fold, include
Proteins Synthesized by Free Ribosomes in the Cytoplasm
So far only proteins synthesized by the ribosomes of the rough E.R. have been
considered. However, most of the proteins of the
mitochondria and
chloroplasts
are synthesized in the in the cytoplasm by
free ribosomes, and are subsequently imported into these organelles, which
requires crossing their membranes. Some mitochondrial proteins are destined for the
matrix, others for the intermembrane space, and others, such as those of
cytochrome c oxidase, for the inner membrane. Some cytoplasmically-synthesized proteins also
end up in peroxisomes.
Most of the mitochondrial proteins produced in the cytoplasm are synthesized as
precursors which have N-terminal signal sequences of
20-60 residues (exceptions include cytochrome c, which is synthesized
in the cytoplasm as the apo- form; after it has entered the mitochondrion,
addition of haem is thought to prevent its transport in the opposite direction).
The uptake of these precursors requires ATP. The conformation of
the soluble precursors would be expected to be different to that of the mature
protein, particularly in the cases of integral membrane proteins.
Some yeast mitochondrial proteins, such as F1-ATPase, are thought to bind in their precursor
form to a receptor on the outer mitochondrial membrane. After the energy-requiring insertion
through the membranes into the matrix, the signal sequence is cleaved (by a metalloprotease).
During or after this cleavage, folding into the mature conformation occurs.
More complicated events accompany the insertion of cytochrome b2 (which
occurs in the intermembrane space of the mitochondria) and cytochrome c1
(which is localized to the outer face of the inner membrane). The precursors of
both of these proteins are partially inserted through the membrane so that a
portion is in the matrix, where a section of the signal sequence is cleaved. The
remaining signal residues are then removed and the haem prosthetic group
is added.
References
- Alberts, B., Bray, D., Lewis, J., Raff, M., Roberts, K. and Watson, J.D.
(1983) Molecular Biology of The Cell, Garland Publishing, New York pp. 340-349
- Darnell, J., Lodish, H. and Baltimore, D. (1986) Molecular Cell Biology,
W.H. Freeman & Co., New York, pp. 940-957
- Stryer, L., (1981) Biochemistry, W.H. Freeman & Co., New York pp. 712-714
- Voet, D. and Voet, J.G. (1990) Biochemistry, John Wiley & Sons, New York
, pp. 298-304
- Walter, P., Gilmore, R. and Blobel, G. (1984) Protein translocation across
the endoplasmic-reticulum Cell 38, 5-8
John Walshaw
 Index to Course Material
Index to Course Material
 Index to Section 4
Index to Section 4
Last updated 11th Feb '96
 Transport of Proteins Across Membranes
Transport of Proteins Across Membranes![]() Index to Course Material
Index to Course Material
![]() Index to Section 4
Index to Section 4
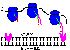 Here is a diagram.
Here is a diagram.