|
Membrane Proteins
Every cell is enclosed by a plasma membrane . In addition, the cytoplasm of eukaryotic cells contains a variety of organelles, each bounded by an internal membrane . All biological membranes are assemblies of lipid and protein molecules held together by non-covalent interactions.
The lipids are amphipathic , ie they have a hydrophilic polar 'head' and a hydrophobic non-polar 'tail', and they therefore spontaneously form bilayers in aqueous solution.
The membrane is fluid, with individual lipid molecules able to diffuse freely within the bilayer. It may therefore be thought of as a '2-dimensional solvent' for the proteins. Proteins diffuse 100 - 100,000 times slower than the lipids. There are three major types of lipids in membranes: phospholipids (the most abundant), glycolipids and cholesterol.
The composition of membranes varies depending on function. The plasma membranes of animal cells are approximately 50% protein in terms of weight. The inner mitochondrial membrane, which is involved in energy transduction, is about 75% protein, while the figure for myelin, which functions as an insulator, is roughly 25%. Many membrane proteins, termed glycoproteins, have covalently bonded carbohydrate groups. In mammalian cells such chains are often attached to an Asparagine's amide group. Carbohydrate chains occur on the extracellular side of plasma membranes, and the luminal side of intracellular membranes around organelles.
Useful links describing membrane structure:
A membrane is not simply a physical barrier between the cell cytoplasm and its external environment, or between different compartments of a cell. The functions of membranes depend on the proteins within them. Some of these proteins have largely a structural role, while many enzymes are associated with membranes.
The manner in which a protein associates with a membrane depends on its structure and can be categorized as follows:
![]() Mellitin
2mlt (46Kb)
[Bbk|BNL|ExP|Waw|Hal]
Use this short script to show the hydrophobic and
polar faces.
Mellitin
2mlt (46Kb)
[Bbk|BNL|ExP|Waw|Hal]
Use this short script to show the hydrophobic and
polar faces.
The region of a protein which traverses the membrane usually consists of an alpha-helix (as in type 2 in the figure) or several alpha-helices ( type 4) each of about 20 residues. In the latter case, the helices are connected by loops which are exposed to the aqueous environment on either side of the membrane and which therefore consist of residues with polar side chains. These connecting loops are generally quite short and may consist of hairpins (see Protein Geometry) .
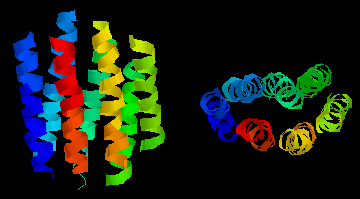 This diagram shows an example of such a "helix-bundle":
bacteriorhodopsin, a light-driven proton-pump from the purple membrane
of Halobacterium halobium . The colours of the 7 helices are simply
to distinguish them in the two perpendicular views. This is a theoretical
model.
This diagram shows an example of such a "helix-bundle":
bacteriorhodopsin, a light-driven proton-pump from the purple membrane
of Halobacterium halobium . The colours of the 7 helices are simply
to distinguish them in the two perpendicular views. This is a theoretical
model.
![]() Click
here 1bac
(119Kb)[Bbk|BNL|ExP|Waw|Hal]
to examine the structure in more detail with RasMol. The 'backbone' or 'ribbons'
options from the 'Display' menu best illustrate the arrangement. The protein
has a bound molecule of retinal which can be seen by selecting 'wireframe'
mode, etc (select the 'chain' option in the 'Colours' menu).
Click
here 1bac
(119Kb)[Bbk|BNL|ExP|Waw|Hal]
to examine the structure in more detail with RasMol. The 'backbone' or 'ribbons'
options from the 'Display' menu best illustrate the arrangement. The protein
has a bound molecule of retinal which can be seen by selecting 'wireframe'
mode, etc (select the 'chain' option in the 'Colours' menu).
![]() This
rhodopsin structure (Halobacterium halobium) was determined by electron
diffraction on two-dimensional crystals
1brd
(124Kb)[Bbk|BNL|ExP|Waw|Hal]
to examine
This
rhodopsin structure (Halobacterium halobium) was determined by electron
diffraction on two-dimensional crystals
1brd
(124Kb)[Bbk|BNL|ExP|Waw|Hal]
to examine
Here is another diagram of bacteriorhodopsin (by Manuel Peitsch at ExPASy). Also study the Bacteriorhodopsin Home Page (by PPS Consultant Iddo Friedberg of the Hebrew University, Jerusalem). If you are able to use MAGE, have a look at the Protein Science kinemages of bacteriorhodopsin modelling and refolding. There is a large super-family of receptors which adopt the same tertiary structure; to learn more about them try the G protein-Coupled Receptor DataBase.
In the case of a single helix, mainly hydrophobic residues would be expected,
so that an apolar surface is exposed to the bilayer. This is illustrated
in the diagram (a) below.
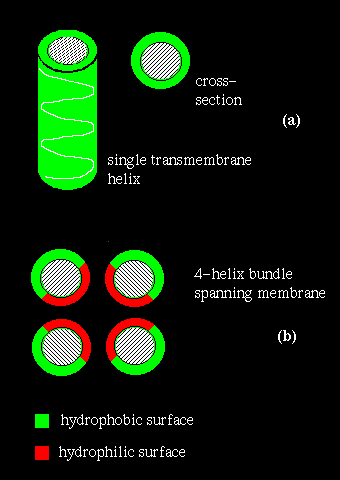
If several helices are bundled, then only the side chains on the outside of the bundle need be hydrophobic. This is illustrated in (b) above. If the central channel between the helices is lined with polar residues, the resulting structure might act as a pore in the membrane through which ions may pass.
Membrane proteins are difficult to crystallize. Because they have significant hydrophilic and hydrophobic surfaces, they are not soluble in aqueous buffer solutions yet they denature in organic solvents. Methods such as Infra-red Spectroscopy, Raman Spectroscopy and Circular Dichroism are used to deduce secondary structure.
However, other approaches may identify transmembrane regions. Naturally, one would expect a relatively long, uninterrupted sequence of hydrophobic residues to represent a membrane-spanning structure. On the other hand, a number of hydrophilic residues would be anticipated in a bundle structure such as the photosynthetic reaction centre (see also previous diagram (b). A quantitative measure of the hydrophobicity of a sequence is required.
Various hydrophobicity scales for amino acids based on for example
the free energy of transfer of the side chain from an inorganic solvent to
water have been compiled. For each position i in the sequence, a
hydropathy index may be calculated as the mean hydrophobicity value
of all the residues from eg i-9 to i+9. Sharp peaks in a plot
of hydropathy index versus residue represent sequences which would be unusually
hydrophobic for a soluble protein and which therefore are strong candidates
for a transmembrane section: 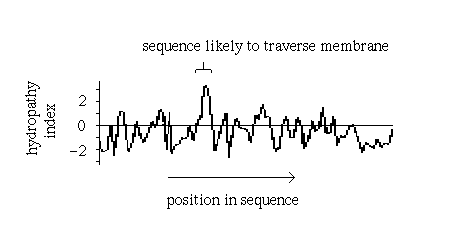
Such plots have in fact been found to be very successful in correctly predicting membrane-spanning sequences in structures which were subsequently elucidated by X-ray crystallography, such as the photosynthetic reaction centre.
The photosynthetic reaction centre of the purple bacterium
Rhodopseudomonas occurs in the membranes of photosynthetic vesicles.
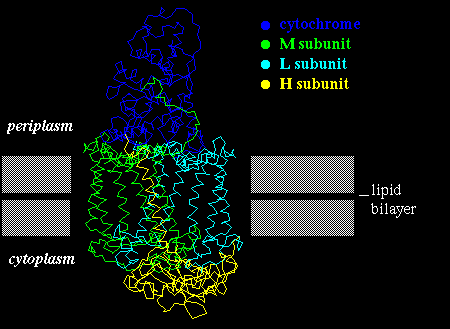
This protein complex is composed of 4 subunits: L, M, H and a
cytochrome. The L and M subunits are homologous and each have 5 transmembrane
alpha-helices. A helix length of 20-25 residues is required to span this
bacterial membrane. The H subunit, on the cytoplasmic side of the membrane,
also has a single transmembrane helix. As in bacteriorhodopsin, the helices
are tilted at an angle of 20-25 ° to the perpendicular to the membrane.
The whole complex is shown below.
![]() Click
here 1prc (910Kb)
[Bbk|BNL|ExP|Waw|Hal]
to look at the crystal structure in more detail.
SCRIPT
Click
here 1prc (910Kb)
[Bbk|BNL|ExP|Waw|Hal]
to look at the crystal structure in more detail.
SCRIPT
A number of pigments (quinones) are bound between the helices of the photosynthetic reaction centre. Buried helix residues which interact either which the pigments, or with other helices are relatively highly conserved between different bacterial species, whereas those on the outside of the bundle are not. This indicates the non-specific nature of the hydrophobic interactions between the complex and the bilayer.
Click here for another diagram. Also look at the following (thanks to Manuel Peitsch at GLAXO Geneva):
Here is the Protein Science kinemage of the photosynthetic reaction centre.
It should be emphasized however that very few high-resolution crystal
structures of membrane proteins exist. Other examples to look at are:
![]() porin 3por
(207Kb)[Bbk|BNL|ExP|Waw|Hal]
(from Rhodobacter ) , which is composed mainly of beta-sheets
in a 16-stranded beta-barrel formation (see section on
all-beta folds ) and forms a pore in the membrane
1.7 - 2.5 nm in diameter (shown below); here are
1,
2
more images of porin.
porin 3por
(207Kb)[Bbk|BNL|ExP|Waw|Hal]
(from Rhodobacter ) , which is composed mainly of beta-sheets
in a 16-stranded beta-barrel formation (see section on
all-beta folds ) and forms a pore in the membrane
1.7 - 2.5 nm in diameter (shown below); here are
1,
2
more images of porin.
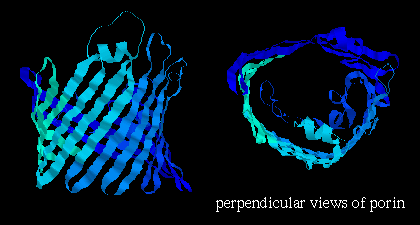
Note that the orientation of the strands is such that side chains alternately point into the interior and exterior of the pore; the former are strongly polar residues while the latter are very hydrophobic. Here is the Protein Science kinemage of the crystal structure. Click here for another kinemage.
![]() Also look at the pore-forming domain of colicin A
1col (260Kb)
[Bbk|BNL|ExP|Waw|Hal].
(There are 2 of these domains in the asymmetric unit of this crystal structure).
Click
here
for a diagram. Colicin A is an antibiotic which is water-soluble, but which
can undergo a conformational change such that the pore-forming domain inserts
itself into the membrane of E. coli .
Also look at the pore-forming domain of colicin A
1col (260Kb)
[Bbk|BNL|ExP|Waw|Hal].
(There are 2 of these domains in the asymmetric unit of this crystal structure).
Click
here
for a diagram. Colicin A is an antibiotic which is water-soluble, but which
can undergo a conformational change such that the pore-forming domain inserts
itself into the membrane of E. coli .
If you are able to use MAGE , click here for the Kinemage supplement to the Branden and Tooze "Introduction to Protein Structure" chapter on membrane proteins.
![]() Also
look at the two kinemages
(1,
2) of
Defensin 1dfn
(53Kb)[Bbk|BNL|ExP|Waw|Hal]
Also
look at the two kinemages
(1,
2) of
Defensin 1dfn
(53Kb)[Bbk|BNL|ExP|Waw|Hal]
|
Last updated 7th April '97