|
|
All-Alpha Topologies
This is not intended to be an exhaustive description of all the known all-alpha folds, but a guide to some of the 'classic' examples.
In addition, it is worthwhile browsing through the Mainly Alpha Class of the CATH Protein Structure Classification Database" at University College, London. This lists 9 orthogonal, and 3 aligned topologies of alpha fold, as well as a solenoid architecture.
You should also be aware of alternative classifications, such as the Structural Classification of Proteins database (Alexey G. Murzin, Steven E. Brenner, Tim J.P. Hubbard, and Cyrus Chothia). In all, seventy-one categories of all alpha folds are listed. A number of the entries have links to diagrams by Manuel Peitsch.
With MAGE installed, study this Kinemage on Alpha Domain Structures, which accompanies the Branden and Tooze book.
There are a number of examples of small proteins (or peptides) which consist of little more than a single helix. A striking example is glucagon, a hormone involved in regulating sugar metabolism in mammals (as is insulin).
![]() Download the glucagon structure: 1gcn (25Kb)
[Bbk|BNL|ExP|Waw|Hal]
Download the glucagon structure: 1gcn (25Kb)
[Bbk|BNL|ExP|Waw|Hal]
Different variations of this supersecondary structure have been introduced
in a previous chapter .
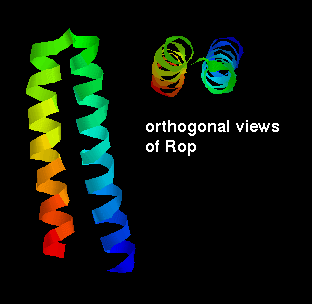
The simplest packing arrangement of a domain of two helices is for them to
lie antiparallel, connected by a short loop. This constitutes the structure
of the small (63 residue) RNA-binding protein Rop, which is found
in certain plasmids (small circular molecules of double-stranded DNA occurring
in bacteria and yeast) and involved in their replication. There is a slight
twist in the arrangement as shown.
![]() 1rop (47Kb)
[Bbk|BNL|ExP|Waw|Hal]
...SCRIPT
1rop (47Kb)
[Bbk|BNL|ExP|Waw|Hal]
...SCRIPT
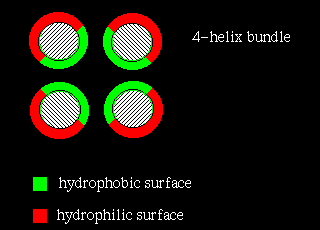
The four-helix bundle is found in a number of different proteins. In many cases the helices part of a single polypeptide chain, connected to each other by three loops. However, the Rop molecule is in fact a dimer of two of the two-helix units shown above.
In four-helix-bundle proteins the interfaces between the helices consist mostly of hydrophobic residues while polar side chains on the exposed surfaces interact with the aqueous environment, as indicated below:
![]() After downloading this ferritin structure, leave the RasMol window
open as we will examine it again shortly.
1fha (121Kb)
[Bbk|BNL|ExP|Waw|Hal]
This SCRIPT colours the side chains of the main
part of the 4 helices (the 4 residues at each end are not shown) as in the
diagram above. 16Kb GIF image.
After downloading this ferritin structure, leave the RasMol window
open as we will examine it again shortly.
1fha (121Kb)
[Bbk|BNL|ExP|Waw|Hal]
This SCRIPT colours the side chains of the main
part of the 4 helices (the 4 residues at each end are not shown) as in the
diagram above. 16Kb GIF image.
Compare this with the arrangement of residues that would be expected in a membrane-spanning helical domain, which is indicated elsewhere in Section 10 (page on membrane proteins) . The central helices of the photosynthetic reaction centre in fact have a similar arrangement to the four- helix bundle.
Other examples exhibit a much more open packing arrangement, as in the
steroid-binding proteins uteroglobin, and
![]() Clara cell 17kDa protein :
1ccd (56Kb)
[Bbk|BNL|ExP|Waw|Hal]
Clara cell 17kDa protein :
1ccd (56Kb)
[Bbk|BNL|ExP|Waw|Hal]
14.6Kb GIF.
The four helices may be arranged in a simple up-and-down topology, as indicated.
 Click here for a diagram
of myohemerythrin, which has this fold.
Click here for a diagram
of myohemerythrin, which has this fold.
![]() To
obtain a similar rendition as in the diagram, use the
colour group and
ribbon commands:
2mhr (201Kb)
[Bbk|BNL|ExP|Waw|Hal]
To
obtain a similar rendition as in the diagram, use the
colour group and
ribbon commands:
2mhr (201Kb)
[Bbk|BNL|ExP|Waw|Hal]
(other examples are cytochrome c':
2ccy (194Kb)
[Bbk|BNL|ExP|Waw|Hal]
and cytochrome b-562: 256b
(174Kb)
[Bbk|BNL|ExP|Waw|Hal]
- which have two molecules in the asymmetric unit; use
restrict *a to show only the A chain).
A more complex arrangement is possible:
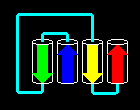 Click here for the
four-helix bundle topology of ferritin.
Click here for the
four-helix bundle topology of ferritin.
![]() If you still
have the RasMol window with the ferritin structure (see above), type:
If you still
have the RasMol window with the ferritin structure (see above), type:
select wireframe off ribbon colour group
Otherwise,
![]() download the structure here:
1fha (121Kb)
[Bbk|BNL|ExP|Waw|Hal]
download the structure here:
1fha (121Kb)
[Bbk|BNL|ExP|Waw|Hal]
The ligand-binding sites of these proteins occur between the alpha helices.
The ligands can be seen by selecting the structures listed above and choosing
the restrict *a,
wireframe and colour chain
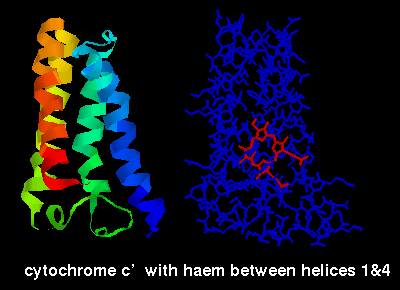
options, e.g. below is a diagram of cytochrome C'
(2ccy- see above for link) indicating the haem
group, which is sited between the first and fourth helices.
A number of cytokines consist of four alpha helices in a bundle. Click here for a diagram of Interleukin-2, human Growth Hormone, Granulocyte-macrophage colony-stimulating factor (GM-CSF) and Interleukin-4, by Manuel Peitsch of Glaxo Geneva. Also examine Simon Brocklehurst's page on cytokines.
DNA-binding domains will be examined in more detail in a later chapter.
Transcription factors are proteins which bind to control regions of DNA. These regions are "upstream" of the structural gene (the sequence which actually codes for a protein) whose transcription they regulate. Transcription factors have a DNA-binding domain and a domain that activates transcription.
The RNA-binding two-helix protein Rop has already been mentioned. A three-helix
bundle forms the basis of a DNA-binding domain which occurs in a number of
proteins- for example homeodomain proteins.
![]() Examine the crystal structure of engrailed homeodomain binding to
DNA, 1hdd (154Kb)
[Bbk|BNL|ExP|Waw|Hal]
and
three
different
diagrams courtesy of Manuel Peitsch.
Examine the crystal structure of engrailed homeodomain binding to
DNA, 1hdd (154Kb)
[Bbk|BNL|ExP|Waw|Hal]
and
three
different
diagrams courtesy of Manuel Peitsch.
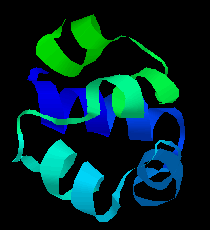
![]() Click here for the structure of the cro repressor from phage 434.
2cro (49Kb)
[Bbk|BNL|ExP|Waw|Hal]
Click here for the structure of the cro repressor from phage 434.
2cro (49Kb)
[Bbk|BNL|ExP|Waw|Hal]
Also refer to the GCN4 transcription factor leucine zipper described in the section on coiled coils, by Antti Iivanainen of the Department of Medical Biochemistry and Biophysics, Karolinska Institute, Stockholm.
The globin fold usually consists of eight alpha helices (e.g. myoglobin). The two helices at the end of the chain are antiparallel, forming a helix-turn-helix motif, but the remainder of the fold does not include any characterized supersecondary structures. These helices pack against each other with larger angles, around 50 °, between them than occurs between antiparallel helices (approximately 20°). See the page in Section 9 on helix-helix packing. Jane Richardson (1981) describes the globin fold as a "Greek key helix bundle", due to the topological similarity with the Greek key arrangement of antiparallel beta-sheets (see section on all beta topologies).
|
|
Last updated 7th April '97