A different helix-loop-helix motif is also common to certain DNA binding proteins. This motif was first observed in prokaryotic DNA binding proteins such as the cro repressor from phage lambda. This protein is a homodimer with each subunit being 66 amino acids in length. Each subunit consists of an all-antiparallel three stranded beta-sheet with three helical segments inserted sequentially between the first and second beta-strands. The two subunits of cro associate by virtue of the third beta-strands which interact forming a six-stranded beta-sheet in the centre of the molecule. Mutagenesis and biochemical work had indicated that residues in the second helix of each cro monomer interacted with DNA. Accordingly model building studies indicated that both these helices in the dimeric protein would fit into the major groove of B-DNA. These proteins recognise base sequences which are palindromic, i.e. possess an internal two-fold symmetry axis. The two recognition helices of the cro protein are also related by a two-fold axis passing through the central beta-sheet region of the dimer. Therefore, the recognition helices of the cro dimer fit into the major groove of the DNA and interact with each identical half of the palindrome. Hence, the second helix of the helix-turn-helix motif has an important role in recognising the DNA while the remainder of the structure serves to keep the two helices in the correct relative position for fitting in the major groove of DNA. Many other helix-turn-helix proteins with different folds exhibit essentially the same mode of binding to DNA.
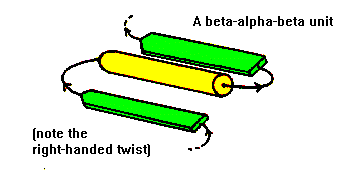
Antiparallel beta-strands can be linked by short lengths of polypeptide forming beta-hairpin structures. In contrast, parallel beta-strands are connected by longer regions of chain which cross the beta-sheet and frequently contain alpha-helical segments. This motif is called the beta-alpha-beta motif and is found in most proteins that have a parallel beta-sheet. The loop regions linking the strands to the helical segments can vary greatly in length. The helix axis is roughly parallel with the beta-strands and all three elements of secondary structure interact forming a hydrophobic core. In certain proteins the loop linking the carboxy terminal end of the first beta-strand to the amino terminal end of the helix is involved in binding of ligands or substrates. The beta-alpha-beta motif almost always has a right-handed fold as demonstrated in the figure.

The subsequent sections of this course on protein folds and domains will demonstrate how the beta-alpha-beta motif is an important and widespread element of supersecondary structure.
Introduction to Protein Structure, C.Branden and J.Tooze, Garland, New York (1991).
A novel super-secondary structure of proteins and the relation between the structure and amino acid sequence, A.F.Efimov, FEBS Lett. 166, 33-38 (1984).
Structure of alpha-alpha-hairpins with short connections, A.V.Efimov, Protein Engin.4, 245-250 (1991).
Structure of beta-beta-hairpins and beta-beta-corners, A.V.Efimov, FEBS Lett., 284, 288-292 (1991).
Structure and evolution of calcium modulated proteins, R.H.Kretsinger, CRC Crit.Rev.Biochem. 8, 119-174 (1980).
Comparison of the structures of cro and lambda repressor proteins in bacteriophage lambda, D.H.Ohlendorf, W.F.Anderson, M.Lewis, C.O.Pabo and B.W.Matthews, J.Molec.Biol. 169 757-769 (1983).
Conformation of beta-hairpins in protein structures, B.L.Sibanda, T.L.Blundell and J.M.Thornton, J.Molec.Biol. 206, 759-777 (1989).
Beta-hairpin families in globular proteins, B.L.Sibanda and J.M.Thornton, Nature 316, 170-174 (1985).