 Thymine and
Guanine
Thymine and
Guanine  Cytosine. Two important properties
of helical dsDNA are the major / minor grooves and the extent
of helical turn or twist.
Cytosine. Two important properties
of helical dsDNA are the major / minor grooves and the extent
of helical turn or twist.
DNA (deoxyribonucleic acid) consists of 4 different bases two
of which are Purines (Adenine and Guanine) and two that are Pyrimidines
(Thymine and Cytosine). DNA exists in both single stranded (ss)
and double stranded (ds) forms but this paper shall only refer
to ds forms. dsDNA can form many different structures. The most
commonly cited structure consists of two anti-parallel sugar-phosphate
backbones forming a right handed helix; the bases from one strand
pointing roughly towards the center of the helix forming a pair
with the base on the opposing strand. The bases are restricted
in their pairing, Adenine 

The two main ds helical forms of DNA are that of A-DNA and B-DNA.
 B-DNA (rasmol
image 37 kb) contains 10 base pairs / 34
B-DNA (rasmol
image 37 kb) contains 10 base pairs / 34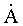 per turn and the 'base pair' plane is at right angles to the long
axis of the helix. This brings about the formation of a Major
and Minor groove with the base pairs placed centrally in the helix.
per turn and the 'base pair' plane is at right angles to the long
axis of the helix. This brings about the formation of a Major
and Minor groove with the base pairs placed centrally in the helix.
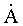 per turn and the 'base pair' plane tilts by about 20o.
This causes the bases to be off center and pushed into the major
groove of the helix, resulting in the minor and major groove being
about equal.
per turn and the 'base pair' plane tilts by about 20o.
This causes the bases to be off center and pushed into the major
groove of the helix, resulting in the minor and major groove being
about equal.
Helical DNA structures are by no means a ridged forming supercoiled
structures, interactions with proteins and other molecules that
result in most 'native' DNA being bent in some way.
Protein recognition of DNA sequence motifs is thought to be a combination of the protein having affinity for DNA with a certain topology (local conformation and configuration) and its ability to form specific H-bonds with the bases in the sequence.
Restriction enzymes differ from gene regulatory proteins being much more fastidious in their substrate choice (a single base pair change eliminates recognition by the protein) and having shorter recognition sequences (4-8 bp vs. 14-20 bp) with strict dyad symmetries. With molecules such as histones or E.coli trp repressor (regulatory proteins) local conformation and configuration of DNA appears to be the only form of recognition.
As noted in the first section DNA can exist in various conformations which would affect its local topology thus either enhance or limit protein recognition. Regulation occurs at the protein level and also at that of DNA conformation. For example if the site of interaction was in the major groove, protein access to the A-DNA would be severely limited compared to if the DNA was in the B form.
Proteins appear to have a diverse but limited number of motifs
that are used in interactions with DNA. These involve both 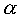
For the sake of comprehension if is easier to think of DNA and
protein in a static form but it is important to remember they
can adopt many conformations, especially on interaction with other
molecules.