 (rasmol image 221 kb)
(rasmol image 221 kb)DNase I (Bovine Pancreatic deoxyribonuclease I) is a 30.4 kDa monomeric glycoprotein that can hydrolyze the P-03'-bond of DNA requiring either Mg2+ and Ca2+ or Mn2+ and Ca2+. The native glycosylated enzyme
 jpg 12 kb. The enzyme is an
jpg 12 kb. The enzyme is an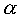 +
+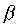 protein (mainly anti-parallel
protein (mainly anti-parallel 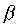 sheet with
segregated
sheet with
segregated 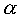 and
and 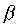 regions). Unlike the majority of characterized DNA binding proteins
it interacts primarily with the minor not the major groove of
DNA and unlike restriction enzymes such as EcoR I its cleavage
is not strictly sequence dependent. Y76 and R41
of DNase I bind within the minor groove (Rasmol script)
but can not form an interaction in A,T rich sequences. This is
thought to be because of the more ridged nature these sequences
impose of the DNA helix.
regions). Unlike the majority of characterized DNA binding proteins
it interacts primarily with the minor not the major groove of
DNA and unlike restriction enzymes such as EcoR I its cleavage
is not strictly sequence dependent. Y76 and R41
of DNase I bind within the minor groove (Rasmol script)
but can not form an interaction in A,T rich sequences. This is
thought to be because of the more ridged nature these sequences
impose of the DNA helix. The binding of the enzyme to DNA causes a distortion of the DNA away from the minor groove, widening it from 11-12
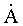 to 15
to 15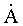 and a bending of the helix axis 21.5o
away from the protein. DNase I binds so that the sugar-phosphate
backbone of the duplex strand to be cleaved is between two
and a bending of the helix axis 21.5o
away from the protein. DNase I binds so that the sugar-phosphate
backbone of the duplex strand to be cleaved is between two 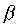 -sheets.
One of these strands makes various interactions with the backbone
of the strand to be cleaved and contains H134 presumed
to act with H252 as general base catalyst essential
for cleavage (to see in rasmol type 'select 134,252','spacefill')
. Another
-sheets.
One of these strands makes various interactions with the backbone
of the strand to be cleaved and contains H134 presumed
to act with H252 as general base catalyst essential
for cleavage (to see in rasmol type 'select 134,252','spacefill')
. Another 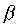 -sheet forms contacts with the
non-cleaved strand and extends on a
-sheet forms contacts with the
non-cleaved strand and extends on a 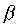 -loop
the two amino acids that bind within the minor groove (Y76
and R41). For DNA cleavage to occur the DNA must be
'bent', therefore sequences that are more ridged (such as A,T
rich regions or DNA that is tightly supercoiled) will be more
resistant to cleavage.
-loop
the two amino acids that bind within the minor groove (Y76
and R41). For DNA cleavage to occur the DNA must be
'bent', therefore sequences that are more ridged (such as A,T
rich regions or DNA that is tightly supercoiled) will be more
resistant to cleavage. (rasmol image of EcoR I monomer and single
strand of the DNA double helix)
(rasmol image of EcoR I monomer and single
strand of the DNA double helix)This is an restriction endonuclease from E.coli that recognizes a strict dyad sequence. EcoR I functions as a dimer of two identical 31 kDa subunits recognizing the 6 bp palindromic sequence G^AATTC with ^ denoting the position of phosphodiester bond cleavage. This DNA sequence imparts a slight curvature that is thought to optimize EcoR I binding. The protein has an
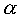 /
/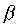 topology mainly parallel
topology mainly parallel 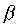 -sheets (
-sheets (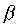 -
-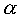 -
-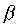 units). Two parallel helices interact directly with the major
groove and are part of a Rossmann (or dinucleotide) fold. See
jpg of EcoR I dimer binding DNA
units). Two parallel helices interact directly with the major
groove and are part of a Rossmann (or dinucleotide) fold. See
jpg of EcoR I dimer binding DNA  (43 kb). It can be seen that it is the N-terminal ends (+ve dipole)
of the helices are brought in contact with the DNA (-ve charge).
(43 kb). It can be seen that it is the N-terminal ends (+ve dipole)
of the helices are brought in contact with the DNA (-ve charge).
Binding of the EcoR I dimer to DNA unwinds it by ~25o (neo-1 kink) widening the major groove by about 3.5
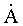 .
This allows the ends of four
.
This allows the ends of four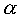 - helices
(two from each subunit) to fit into the major groove. This widening
of the major groove also results in numerous H-bonds between the
DNA and protein (R 145,200 and E 144 of
each subunit) that are not possible undistorted DNA (rasmol script).
The specific H-bonds are R200 to N7 and
O9 of guanine while the two adjacent adenines are cross
linked by R145 donating H-bonds to N7 of
the two adjacent base pairs and E144 from the other
subunit accepting H-bonds from N6 of the same adenines.
The result is two helices from each subunit are pushed into the
major groove forming specific interactions. The cross-linking
interactions of R145 and E144 are thought
to be especially important in imparting the specific nature of
the interaction. It is apparent the formation of neo-1 kinks at
random to make the specific interactions would be very energetically
unfavorable. It is therefore presumed that the intrinsic topology
of the recognition sequence or similar sequences must be 'recognized'
by the enzyme before forming the neo-1 kink. Specific binding
of EcoR I to DNA also brings about kinks in the DNA at the ends
of the recognition sequence (neo-2 kink).
- helices
(two from each subunit) to fit into the major groove. This widening
of the major groove also results in numerous H-bonds between the
DNA and protein (R 145,200 and E 144 of
each subunit) that are not possible undistorted DNA (rasmol script).
The specific H-bonds are R200 to N7 and
O9 of guanine while the two adjacent adenines are cross
linked by R145 donating H-bonds to N7 of
the two adjacent base pairs and E144 from the other
subunit accepting H-bonds from N6 of the same adenines.
The result is two helices from each subunit are pushed into the
major groove forming specific interactions. The cross-linking
interactions of R145 and E144 are thought
to be especially important in imparting the specific nature of
the interaction. It is apparent the formation of neo-1 kinks at
random to make the specific interactions would be very energetically
unfavorable. It is therefore presumed that the intrinsic topology
of the recognition sequence or similar sequences must be 'recognized'
by the enzyme before forming the neo-1 kink. Specific binding
of EcoR I to DNA also brings about kinks in the DNA at the ends
of the recognition sequence (neo-2 kink). It appears that the binding of EcoR I to DNA and subsequent cleavage are to distinct activities of EcoR I. For example EcoR I can bind to DNA in the absence of Mg2+ but cleavage is dependent on the Mg2+ being complexed; mutation studies have shown that replacement of E111 (well away from the protein-DNA interface) with Gly results in an protein still capable of binding DNA but not of cleavage.