 PPS96 Projects
PPS96 Projects  PPS96 Projects
PPS96 Projects
Cristina Cantale
Integration mechanism
Since then, a large amount of work has been carried out using in vitro
systems and mainly HIV IN purified protein (but also INs from ASLV, RSV (Rous
Sarcoma virus), MoMULV (Moloney Murine leukemia virus) and very recently HTLV-II
(Human T-cell Leukemia virus type II)), with the aim of clarifying the
integration mechanism.
It was demonstrated that IN is able to carry out
three different reactions:
The overall reaction carried out by IN is a transesterification, produced by
a nucleophilic attack on an activated phosphodiester bond performed by water or
by the recessed 5'CA(OH)3' hydroxyl group end.
It appears that the same
catalytic core domain is involved in both processing and the DNA transfer.
Both
reactions proceed by a one step mechanism, as demonstrated using a known
chirality substrate (phosphorothioate), without the formation of covalent
protein-DNA intermediate
(Engelman A. et al., 91)
These reactions do not need any
external energy but a divalent cation (Mg2+ or Mn2+) is
necessary for the reaction to proceed. Beside the hypotheses of involvement of
divalent cations in active core mechanism, a novel in vitro assay using
immobilized LTR oligonucleotides (Hazuda et al., 94-1) suggests
that the requirement for Mn2+ is correlated with the formation of
the oligomeric structure of IN in solution (Wolfe et al., 96).
This is considered the very first step of the overall integration reaction, that
is the assembly of a stable complex between integrase and viral DNA
(Ellison V. and Brown P.O., 94 - Ellison V. et al., 95
-
Vink C. et al., 94).
As two sterically and temporally
coordinated reactions (one at each end of viral DNA) are required for
integration of viral DNA, IN has to be at least a dimer, carrying the double
strand viral DNA.
Staggered cleavage of the host DNA should involve another
dimer (at least), suggesting that IN works as a tetramer (at least).
Complementation experiments using IN mutants lacking different portions support
this hypothesis
(van Gent et al., 93 - Jones C.S. et al., 92 -
Engelman A. et al.,93).
The use of mutants has proved also
that there are different domains of IN which play different roles
(transesterification, multimerization, DNA recognition).Furthermore it has
permitted the identification of the AAs that are fundamental for IN activities,
confirming the results obtained from the sequence analysis of various retroviral
INs.
To complete the picture, it should be emphasized that, even if the
in vitro experiments have the great worth to have clarified many aspects of
IN behaviour, they are not able to simulate entirely the in vivo system.
The
same mechanism of integration is only partially reproduced.
The in vitro
system lacks the aspects of concerted two ends strand transfer reactions and
just one strand is processed and joined to the DNA target, with the final
product having a typical Y form.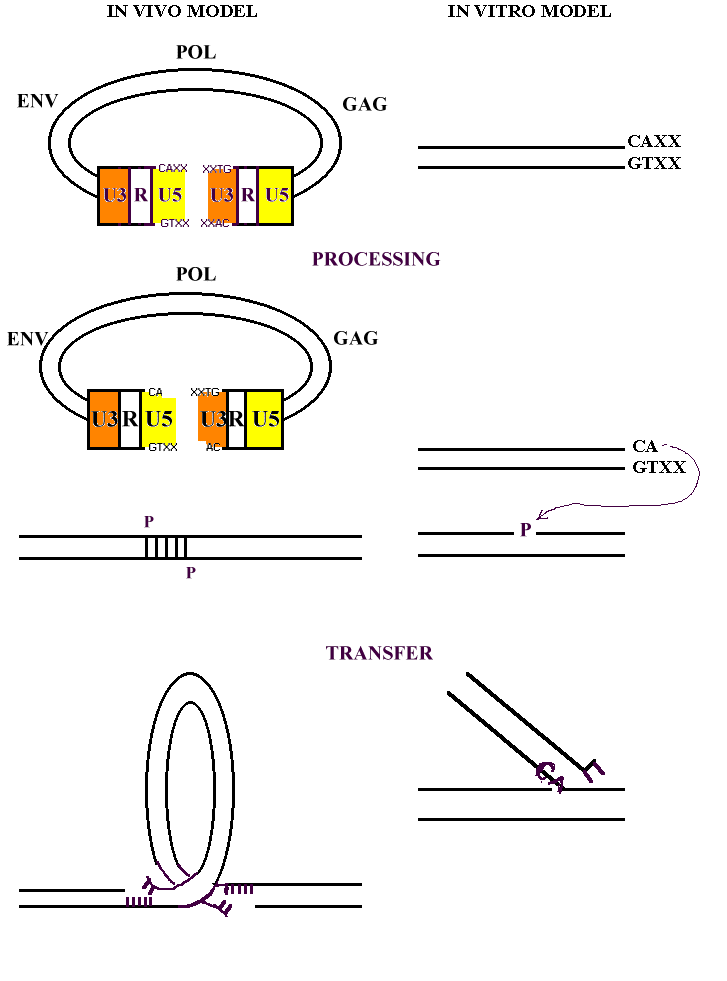
Along with the actual mechanism, the aspects connected with viral DNA and
host DNA recognition characteristics need to be deepened.
As
previously reported, the specificity of viral
DNA for LTR sequences is not so high and there are reported examples of IN
proteins able to react with an oligonucleotide simulating LTR from different
retroviruses (e.g. MoMULV IN with HIV LTR ends in an aspecific fashion, but not
the reverse (Vink C. et al., 91-2)). Nucleotides next to the
subterminal CA have been reported to be involved, namely the subterminal 6 to 8
nucleotides
(Reicin A.S. et al., 95). The prevailing idea is that the
specificity is not mainly connected with sequence but with other aspects of LTR
viral DNA.
Hovewer it has been underlined that in the in vivo
systems IN and the viral DNA are not free in cytoplasm, but both are part of an
ordered complex, the PreIntegration Complex (PIC). PICs are so stable
assemblages that they can be extracted from cytoplasm of infected cells retainig
their activities.
Consequentely, it has been proposed that IN doesn't need
such a large sequence specificity to recognize its substrate and only a short
repeat CA, highly conserved, is essential for right positioning and catalysis
(van Gent et al., 91 - Hazuda et al., 94-2) ,
together with a subterminal portion interacting with the HHCC region of IN
(Vincent K.A. et al., 93).
The main aspects promoting IN
attack on host DNA for strand transfer reaction are not still completely
understood.
It seems that in vivo the site of attack is strongly
influenced by chromatin. There are some preferences, like regions complexed
with transcription factors (Kassavetis et al., 89) or by histones
(Morse et al., 92) or DNaseI sensitive sites. Probably there is
some sequence bias, too.
Some in vitro experiments were carried out using
more and more complex target DNA structures; a particularly efficient
integration into nucleosomal DNA
(Pryciak P.M. and Varmus E.H., 92) and in the most severely deformed and
kinked DNA regions within the nucleosomal core (Pruss et al., 94),
was observed.
It has been proposed that this is due to the bending of DNA in
these regions, which may activate integration (Muller H.P. et al., 94).The
bending promotes a DNA conformation (which is favourable for integration),
widening the minor and/or major groove(s) on the exposed face of the DNA helix.
There are also other parameters that can be influenced by DNA bending, like
affinity for Mn 2+; also transfer reaction might require local
denaturation of DNA, easier in a bent region.
In any case, the in vivo
system is very complex: specific interations with host proteins have to be
taken into account, following what observed for retrotransposone Ty3 (Chalker
D.L. and Sandmeyer S.B., 92), together with the subnuclear localization
of viral PIC; the host cell state during integration could play a role too. Such
interactions have been also proposed to explain the capacity of retroviral DNA
to protect itself from the autointegration process (Lee M.S. and Craige
R., 94).
![]() PPS96
PPS96![]() List of Contents
List of Contents![]() A
bit of history
A
bit of history![]() The Sequence
The Sequence![]() References
References
Last updated 25th Oct '96