 Beta-strand Packing in Alpha/Beta Barrels
Beta-strand Packing in Alpha/Beta Barrels
![]() Index to Course Material
Index to Course Material
![]() Index to Section 9
Index to Section 9
![]() Sheet-Sheet Packing
Sheet-Sheet Packing
![]() Helix-Sheet Packing
Helix-Sheet Packing
In the classic form of the barrel, a three-residue section of each beta-strand contributes to the core. Referring to these residues as i, i+1 and i+2, alternate beta-strands have 2 (i, i+2) and 1 (i+1) side chains in the core, which therefore consists of 3 layers of 4 side chains each.
![]() This is illustrated in the structure of glycolate oxidase, 1gox (259Kb)
[Bbk|BNL|ExP|Waw|Hal]
by this SCRIPT.
This is illustrated in the structure of glycolate oxidase, 1gox (259Kb)
[Bbk|BNL|ExP|Waw|Hal]
by this SCRIPT.
This is also shown in this diagram (15Kb GIF), showing a top and side view.
The residues in each of the three layers are coloured blue, green and red. The orientations of each side chain are indicated. The RasMol script defines the layers as the sets top, middle and bottom, and all three collectively as core. After select-ing one of the sets, use spacefill to display the sidechains. For example to display the whole core, cut-and-paste the following:
select core spacefill select *.CA colour yellowThe result is shown in this diagram (17Kb GIF).
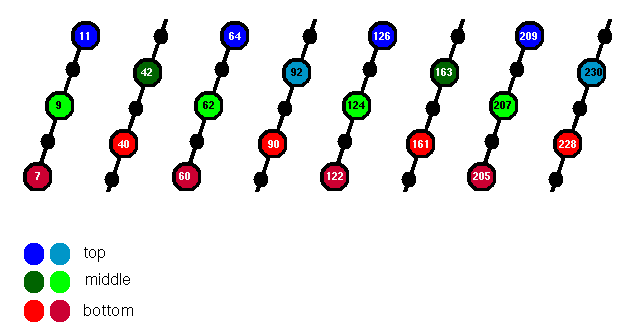
Figure 3. Five residues of each beta-strand of the TIM barrel are indicated; those whose side chains point into the core are highlighted. The top, middle and bottom layers are coloured blue, green and red respectively.
![]() Download the triose phosphate isomerase structure, 1tim (322Kb)
[Bbk|BNL|ExP|Waw|Hal]
and then use this RasMol SCRIPT. It defines various
sets which will be used below. The result should appear as this
diagram (18Kb GIF).
Download the triose phosphate isomerase structure, 1tim (322Kb)
[Bbk|BNL|ExP|Waw|Hal]
and then use this RasMol SCRIPT. It defines various
sets which will be used below. The result should appear as this
diagram (18Kb GIF).
Designating the 5 residues of each strand as i ... i+4, then alternate strands contribute the side chains of i and i+1 to the core. The two resulting sub-layers are coloured different shades of red, but it is apparent that they intercalate fairly efficiently to form a single layer, with the exception of the polar Arg 205 which is positioned underneath.
To display this layer, type:
select bottom spacefillN.B. the sub-layers are also defined, named bottom1 and bottom2 respectively.
Alternate strands have the side chain of i+2 in the core (this sub-layer is called middle1). Two i+3 side chains (middle2) are at approximately the same level. To display the whole middle section:
select middle spacefillThe top layer is similar to the middle (inverted), consisting of two opposing i+3 side chains (top1) and the four i+4 side chains of alternate strands (top2).
select top spacefillNote that there is considerable intercalation between the middle and top layers, so that this classification is somewhat arbitrary. The middle2 and top1 side chains are at approximately the same level. Several of the core residues are glycines, but the main chain atoms contribute.
To highlight the orientation of the side chains pointing away from the core:
select outer wireframe 40
Last updated 1st Aug '96