 Serine Proteases Part 3
Serine Proteases Part 3
![]() Index to Course Material
Index to Course Material
![]() Index to Section 12
Index to Section 12
![]() Enzymes Index;
Enzymes Index;
![]() Part 2
Part 2
![]() Part 4
Part 4
![]() 8gch (185Kb)
[Bbk|BNL|ExP|Waw|Hal]
8gch (185Kb)
[Bbk|BNL|ExP|Waw|Hal]
The active site lies between the two domains. It consists of residues on the same two loops (firstly between beta-strands 3 and 4, secondly between beta strands 5 and 6) of each of the two barrel domains. Four features of the active site are now described.
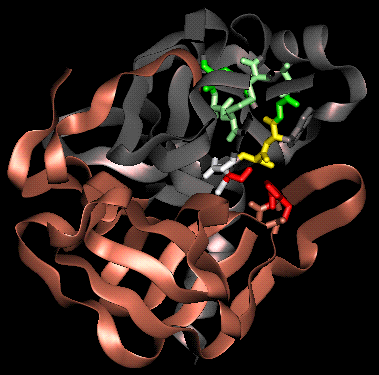
(refer to text for colour coding)
SCRIPT 1 renders the 8gch structure as in the diagram.
 The Catalytic Triad
The Catalytic TriadThe catalytic triad has been introduced in page 1. This introduced the charge relay network model, in which His 57 is polarized by nearby Asp 102, promoting its capacity as a proton shuttle: when the Ser-195 O-gamma (OG) attacks the substrate, the serine hydrogen is transferred to His 57. The nucleophilicity of Ser-195 is increased by the histidine side chain.
Initially, a proton was thought to be transferred from His 57 to Asp 102; in this scenario the charge on the Asp side chain would therefore be relayed to the serine O-gamma during catalysis, since the latter's hydrogen ion is transferred to the His.
However, subsequent neutron diffraction (Kossiakoff and Spencer,1981) and N15-NMR studies (Bachovchin, 1985) indicated that in the tetrahedral intermediate, the proton remains bound to N-delta-1 (ND1) of the imidazole ring. This means that the ring becomes ionized (imidazolium ion), as it accepts a proton from the serine but does not donate one to the aspartate.
The function of Asp-102 is thought to involve polarization of the His side chain, and stabilization of the necessary protonation state of the imidazole ring, i.e. with the hydrogen situated on the N-delta-1 atom rather than on N-epsilon-2 (NE2), so that that latter is able to accept the proton from OG. The Asp stabilizes the positively-charged imidazolium ion which results, and maintains the His side chain in the appropriate conformation.
 The Oxyanion Hole
The Oxyanion HoleX-ray crystal structures and chemical data led to the proposal of a tetrahedral transition-state intermediate in the catalytic mechanism of chymotrypsin. This is formed by the attack of the Ser-195 on the carbonyl carbon atom of the scissile bond. The C=O bond becomes a single bond, leaving a negative charge on the O atom (an oxyanion), while the fourth valency of the carbon atom is occupied by a bond with the serine O-gamma. The oxyanion forms hydrogen bonds to two main chain amides, of residues 193 (Gly) and 195 (Ser). This binding site is termed the oxyanion hole.
Note that the active site has a complementary structure to the transition state.
 The Substrate Specificity Pocket
The Substrate Specificity PocketThe substrate specificity pocket accommodates the side chain of the residue preceding the scissile bond, and has already been briefly described in page 1. Principal side chains lining the pocket are those of residues 189,216 and 226, since they are found to be different in trypsin, chymotrypsin and elastase. 189 is at the bottom of the pocket, while 216 and 226 are situated on the sides.
(Residues 189, 216 and 226 in  bright
green; other residues of the pocket highlighted in
bright
green; other residues of the pocket highlighted in
 duller green)
duller green)
 Main Chain Substrate-binding
Main Chain Substrate-bindingApart from the specificity pocket, the substrate binding site is not in the form of a groove or cleft, but runs across the surface of the enzyme. The substrate hydrogen bonds to a short region (residues 214 to 216) of the enzyme main chain in an antiparallel beta-sheet arrangement. The subsites which bind the three substrate residues preceding the scissile bond are termed S1, S2 and S3 (S1, the primary binding site, is the specificity pocket; subsites on the other side of the scissile bond are named S1', S2' etc).
(Note that residue 216 is couloured green in the diagrams as it is an important feature of the specificity pocket)
Last updated 11th Jul '96