 Equine Alcohol Dehydrogenase
Equine Alcohol Dehydrogenase
![]() Index to Course Material
Index to Course Material
![]() Index to Section 11
Index to Section 11
![]() Multimeric Enzymes
Multimeric Enzymes
Each of the two subunits of the horse enzyme has one binding site for NAD+ and two binding sites for Zn2+. Only one of the zinc ions is involved directly in catalysis. It is ligated by the side chains of Cys 46, His 67, Cys 174 and a water molecule which hydrogen bonds to Ser 48. (If you wish to view this water molecule, download the 8adh PDB structure, which is a monomer without NAD+, as the 6adh dimer includes no water molecules.)
The zinc ion is situated at the junction of two binding sites: one of these is a pocket which binds NAD+, while the other is a cleft which binds the substrate (which is one of a variety of aromatic and aliphatic alcohols).
![]() Download one or both of these crystal structures of horse liver alcohol
dehydrogenase:
Download one or both of these crystal structures of horse liver alcohol
dehydrogenase:
 Click here for a diagram
of the dimer; use this SCRIPT 2 on the 6adh
structure (see above) to render as in this diagram
Click here for a diagram
of the dimer; use this SCRIPT 2 on the 6adh
structure (see above) to render as in this diagram
 and here for a space-filling
model, where the NAD+ molecules are visible. If you have just run
SCRIPT 2, type rotate y 180 followed by
spacefill in the RasMol command line window to obtain this rendition
and here for a space-filling
model, where the NAD+ molecules are visible. If you have just run
SCRIPT 2, type rotate y 180 followed by
spacefill in the RasMol command line window to obtain this rendition
The domain involved in subunit-subunit interactions includes a six-stranded parallel beta sheet- this is a doubly wound alpha/beta fold (link to Section 10). The strands of each subunit are in an antiparallel alignment with respect to each other, giving a twelve-stranded mixed beta sheet.
 This is indicated in this
diagram . Obtain a similar rendition with
SCRIPT 3.
This is indicated in this
diagram . Obtain a similar rendition with
SCRIPT 3.
In order to visualize how two monomers pack together to form a dimer, we will concentrate on the positions of the side chains of His-105 and Tyr-286. The face of the histidine of one subunit stacks against the tyrosine of the other, and vice-versa. To highlight these, type the RasMol commands:
select (his105.CA,tyr286.CA),(his105,tyr286) and sidechain wireframe 80
 This diagram illustrates
this packing. Or use this short SCRIPT 4.
This diagram illustrates
this packing. Or use this short SCRIPT 4.
The following diagram indicates the positions of these side chains in relation to the interfacial beta strands and the monomer as a whole.
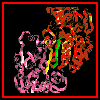 The symmetry of the monomer-
monomer interaction is shown here.
SCRIPT 6
The symmetry of the monomer-
monomer interaction is shown here.
SCRIPT 6
A larger scale diagram of the dimer is shown. Make sure you can locate the axis of symmetry.
Consider how precise the interaction surface of the monomer must be in order for this dimerization to occur.
Note that in other organisms the enzyme exists in different multimeric forms. For example, the subunit of yeast alcohol dehydrogenase has a high degree of homology with the horse liver protein, but is tetrameric.
Last updated 28th Jun '96